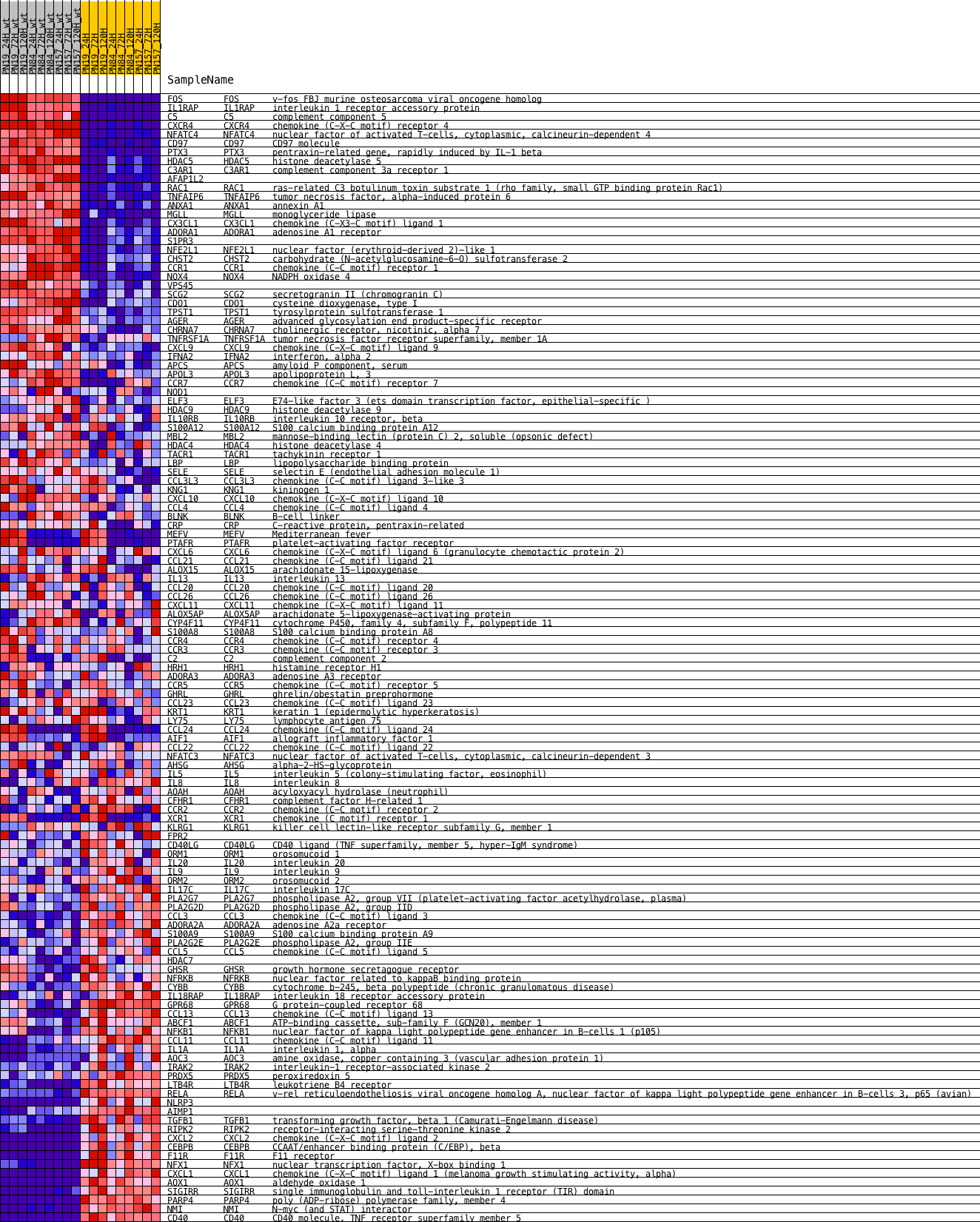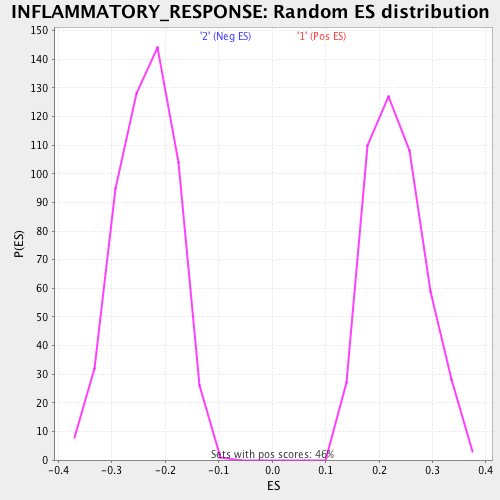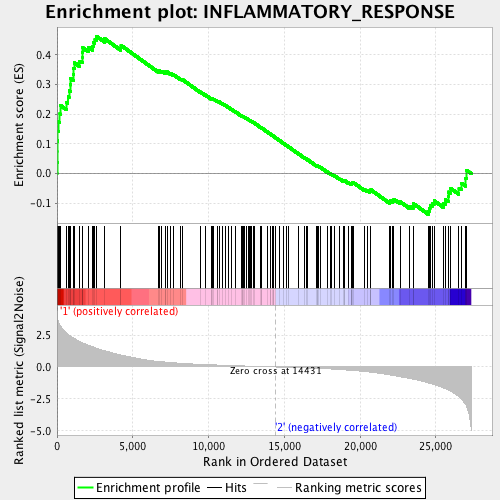
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | expression_mean_normalized_with_dChip_geneAvg_Mar1014_PNs.mainSample.cls #wt_versus_radiation.mainSample.cls #wt_versus_radiation_repos |
| Phenotype | mainSample.cls#wt_versus_radiation_repos |
| Upregulated in class | 1 |
| GeneSet | INFLAMMATORY_RESPONSE |
| Enrichment Score (ES) | 0.46309412 |
| Normalized Enrichment Score (NES) | 2.0049794 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0033325064 |
| FWER p-Value | 0.008 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FOS | NA | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 | 4.177 | 0.0384 | Yes |
| 2 | IL1RAP | NA | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 13 | 3.933 | 0.0743 | Yes |
| 3 | C5 | NA | C5 Entrez, Source | complement component 5 | 15 | 3.923 | 0.1105 | Yes |
| 4 | CXCR4 | NA | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 51 | 3.653 | 0.1429 | Yes |
| 5 | NFATC4 | NA | NFATC4 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | 78 | 3.543 | 0.1746 | Yes |
| 6 | CD97 | NA | CD97 Entrez, Source | CD97 molecule | 187 | 3.291 | 0.2010 | Yes |
| 7 | PTX3 | NA | PTX3 Entrez, Source | pentraxin-related gene, rapidly induced by IL-1 beta | 227 | 3.226 | 0.2293 | Yes |
| 8 | HDAC5 | NA | HDAC5 Entrez, Source | histone deacetylase 5 | 620 | 2.653 | 0.2394 | Yes |
| 9 | C3AR1 | NA | C3AR1 Entrez, Source | complement component 3a receptor 1 | 718 | 2.553 | 0.2593 | Yes |
| 10 | AFAP1L2 | NA | 790 | 2.486 | 0.2796 | Yes | ||
| 11 | RAC1 | NA | RAC1 Entrez, Source | ras-related C3 botulinum toxin substrate 1 (rho family, small GTP binding protein Rac1) | 869 | 2.415 | 0.2990 | Yes |
| 12 | TNFAIP6 | NA | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 885 | 2.401 | 0.3206 | Yes |
| 13 | ANXA1 | NA | ANXA1 Entrez, Source | annexin A1 | 1067 | 2.267 | 0.3349 | Yes |
| 14 | MGLL | NA | MGLL Entrez, Source | monoglyceride lipase | 1086 | 2.251 | 0.3550 | Yes |
| 15 | CX3CL1 | NA | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 1151 | 2.210 | 0.3730 | Yes |
| 16 | ADORA1 | NA | ADORA1 Entrez, Source | adenosine A1 receptor | 1490 | 1.984 | 0.3789 | Yes |
| 17 | S1PR3 | NA | 1659 | 1.893 | 0.3902 | Yes | ||
| 18 | NFE2L1 | NA | NFE2L1 Entrez, Source | nuclear factor (erythroid-derived 2)-like 1 | 1666 | 1.889 | 0.4074 | Yes |
| 19 | CHST2 | NA | CHST2 Entrez, Source | carbohydrate (N-acetylglucosamine-6-O) sulfotransferase 2 | 1693 | 1.877 | 0.4237 | Yes |
| 20 | CCR1 | NA | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 2059 | 1.692 | 0.4259 | Yes |
| 21 | NOX4 | NA | NOX4 Entrez, Source | NADPH oxidase 4 | 2355 | 1.572 | 0.4296 | Yes |
| 22 | VPS45 | NA | 2377 | 1.560 | 0.4432 | Yes | ||
| 23 | SCG2 | NA | SCG2 Entrez, Source | secretogranin II (chromogranin C) | 2497 | 1.506 | 0.4527 | Yes |
| 24 | CDO1 | NA | CDO1 Entrez, Source | cysteine dioxygenase, type I | 2583 | 1.467 | 0.4631 | Yes |
| 25 | TPST1 | NA | TPST1 Entrez, Source | tyrosylprotein sulfotransferase 1 | 3140 | 1.262 | 0.4543 | No |
| 26 | AGER | NA | AGER Entrez, Source | advanced glycosylation end product-specific receptor | 4213 | 0.928 | 0.4234 | No |
| 27 | CHRNA7 | NA | CHRNA7 Entrez, Source | cholinergic receptor, nicotinic, alpha 7 | 4214 | 0.927 | 0.4320 | No |
| 28 | TNFRSF1A | NA | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 6672 | 0.432 | 0.3457 | No |
| 29 | CXCL9 | NA | CXCL9 Entrez, Source | chemokine (C-X-C motif) ligand 9 | 6726 | 0.426 | 0.3476 | No |
| 30 | IFNA2 | NA | IFNA2 Entrez, Source | interferon, alpha 2 | 6917 | 0.405 | 0.3444 | No |
| 31 | APCS | NA | APCS Entrez, Source | amyloid P component, serum | 7147 | 0.382 | 0.3395 | No |
| 32 | APOL3 | NA | APOL3 Entrez, Source | apolipoprotein L, 3 | 7159 | 0.381 | 0.3426 | No |
| 33 | CCR7 | NA | CCR7 Entrez, Source | chemokine (C-C motif) receptor 7 | 7252 | 0.370 | 0.3426 | No |
| 34 | NOD1 | NA | 7478 | 0.346 | 0.3375 | No | ||
| 35 | ELF3 | NA | ELF3 Entrez, Source | E74-like factor 3 (ets domain transcription factor, epithelial-specific ) | 7679 | 0.327 | 0.3332 | No |
| 36 | HDAC9 | NA | HDAC9 Entrez, Source | histone deacetylase 9 | 8162 | 0.287 | 0.3181 | No |
| 37 | IL10RB | NA | IL10RB Entrez, Source | interleukin 10 receptor, beta | 8274 | 0.280 | 0.3166 | No |
| 38 | S100A12 | NA | S100A12 Entrez, Source | S100 calcium binding protein A12 | 9458 | 0.206 | 0.2750 | No |
| 39 | MBL2 | NA | MBL2 Entrez, Source | mannose-binding lectin (protein C) 2, soluble (opsonic defect) | 9779 | 0.189 | 0.2650 | No |
| 40 | HDAC4 | NA | HDAC4 Entrez, Source | histone deacetylase 4 | 10190 | 0.170 | 0.2515 | No |
| 41 | TACR1 | NA | TACR1 Entrez, Source | tachykinin receptor 1 | 10198 | 0.170 | 0.2528 | No |
| 42 | LBP | NA | LBP Entrez, Source | lipopolysaccharide binding protein | 10223 | 0.169 | 0.2535 | No |
| 43 | SELE | NA | SELE Entrez, Source | selectin E (endothelial adhesion molecule 1) | 10339 | 0.163 | 0.2508 | No |
| 44 | CCL3L3 | NA | CCL3L3 Entrez, Source | chemokine (C-C motif) ligand 3-like 3 | 10566 | 0.152 | 0.2439 | No |
| 45 | KNG1 | NA | KNG1 Entrez, Source | kininogen 1 | 10587 | 0.152 | 0.2446 | No |
| 46 | CXCL10 | NA | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 10692 | 0.147 | 0.2421 | No |
| 47 | CCL4 | NA | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 10911 | 0.137 | 0.2353 | No |
| 48 | BLNK | NA | BLNK Entrez, Source | B-cell linker | 11083 | 0.128 | 0.2302 | No |
| 49 | CRP | NA | CRP Entrez, Source | C-reactive protein, pentraxin-related | 11308 | 0.117 | 0.2231 | No |
| 50 | MEFV | NA | MEFV Entrez, Source | Mediterranean fever | 11512 | 0.110 | 0.2166 | No |
| 51 | PTAFR | NA | PTAFR Entrez, Source | platelet-activating factor receptor | 11738 | 0.100 | 0.2093 | No |
| 52 | CXCL6 | NA | CXCL6 Entrez, Source | chemokine (C-X-C motif) ligand 6 (granulocyte chemotactic protein 2) | 12156 | 0.084 | 0.1947 | No |
| 53 | CCL21 | NA | CCL21 Entrez, Source | chemokine (C-C motif) ligand 21 | 12242 | 0.081 | 0.1924 | No |
| 54 | ALOX15 | NA | ALOX15 Entrez, Source | arachidonate 15-lipoxygenase | 12260 | 0.080 | 0.1925 | No |
| 55 | IL13 | NA | IL13 Entrez, Source | interleukin 13 | 12270 | 0.080 | 0.1929 | No |
| 56 | CCL20 | NA | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 12374 | 0.076 | 0.1898 | No |
| 57 | CCL26 | NA | CCL26 Entrez, Source | chemokine (C-C motif) ligand 26 | 12485 | 0.072 | 0.1864 | No |
| 58 | CXCL11 | NA | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 12633 | 0.066 | 0.1816 | No |
| 59 | ALOX5AP | NA | ALOX5AP Entrez, Source | arachidonate 5-lipoxygenase-activating protein | 12698 | 0.064 | 0.1799 | No |
| 60 | CYP4F11 | NA | CYP4F11 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 11 | 12758 | 0.062 | 0.1783 | No |
| 61 | S100A8 | NA | S100A8 Entrez, Source | S100 calcium binding protein A8 | 12854 | 0.059 | 0.1753 | No |
| 62 | CCR4 | NA | CCR4 Entrez, Source | chemokine (C-C motif) receptor 4 | 12958 | 0.054 | 0.1720 | No |
| 63 | CCR3 | NA | CCR3 Entrez, Source | chemokine (C-C motif) receptor 3 | 13040 | 0.051 | 0.1695 | No |
| 64 | C2 | NA | C2 Entrez, Source | complement component 2 | 13410 | 0.037 | 0.1563 | No |
| 65 | HRH1 | NA | HRH1 Entrez, Source | histamine receptor H1 | 13455 | 0.035 | 0.1550 | No |
| 66 | ADORA3 | NA | ADORA3 Entrez, Source | adenosine A3 receptor | 13509 | 0.033 | 0.1534 | No |
| 67 | CCR5 | NA | CCR5 Entrez, Source | chemokine (C-C motif) receptor 5 | 13865 | 0.020 | 0.1405 | No |
| 68 | GHRL | NA | GHRL Entrez, Source | ghrelin/obestatin preprohormone | 14071 | 0.012 | 0.1331 | No |
| 69 | CCL23 | NA | CCL23 Entrez, Source | chemokine (C-C motif) ligand 23 | 14077 | 0.012 | 0.1330 | No |
| 70 | KRT1 | NA | KRT1 Entrez, Source | keratin 1 (epidermolytic hyperkeratosis) | 14206 | 0.008 | 0.1284 | No |
| 71 | LY75 | NA | LY75 Entrez, Source | lymphocyte antigen 75 | 14245 | 0.006 | 0.1270 | No |
| 72 | CCL24 | NA | CCL24 Entrez, Source | chemokine (C-C motif) ligand 24 | 14275 | 0.005 | 0.1260 | No |
| 73 | AIF1 | NA | AIF1 Entrez, Source | allograft inflammatory factor 1 | 14416 | 0.001 | 0.1209 | No |
| 74 | CCL22 | NA | CCL22 Entrez, Source | chemokine (C-C motif) ligand 22 | 14698 | -0.009 | 0.1106 | No |
| 75 | NFATC3 | NA | NFATC3 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 3 | 14968 | -0.020 | 0.1009 | No |
| 76 | AHSG | NA | AHSG Entrez, Source | alpha-2-HS-glycoprotein | 15122 | -0.026 | 0.0955 | No |
| 77 | IL5 | NA | IL5 Entrez, Source | interleukin 5 (colony-stimulating factor, eosinophil) | 15268 | -0.032 | 0.0905 | No |
| 78 | IL8 | NA | IL8 Entrez, Source | interleukin 8 | 15907 | -0.056 | 0.0675 | No |
| 79 | AOAH | NA | AOAH Entrez, Source | acyloxyacyl hydrolase (neutrophil) | 16347 | -0.075 | 0.0521 | No |
| 80 | CFHR1 | NA | CFHR1 Entrez, Source | complement factor H-related 1 | 16434 | -0.079 | 0.0497 | No |
| 81 | CCR2 | NA | CCR2 Entrez, Source | chemokine (C-C motif) receptor 2 | 16547 | -0.083 | 0.0463 | No |
| 82 | XCR1 | NA | XCR1 Entrez, Source | chemokine (C motif) receptor 1 | 17115 | -0.106 | 0.0264 | No |
| 83 | KLRG1 | NA | KLRG1 Entrez, Source | killer cell lectin-like receptor subfamily G, member 1 | 17150 | -0.108 | 0.0262 | No |
| 84 | FPR2 | NA | 17173 | -0.109 | 0.0264 | No | ||
| 85 | CD40LG | NA | CD40LG Entrez, Source | CD40 ligand (TNF superfamily, member 5, hyper-IgM syndrome) | 17229 | -0.112 | 0.0254 | No |
| 86 | ORM1 | NA | ORM1 Entrez, Source | orosomucoid 1 | 17369 | -0.118 | 0.0214 | No |
| 87 | IL20 | NA | IL20 Entrez, Source | interleukin 20 | 17826 | -0.143 | 0.0059 | No |
| 88 | IL9 | NA | IL9 Entrez, Source | interleukin 9 | 18039 | -0.155 | -0.0004 | No |
| 89 | ORM2 | NA | ORM2 Entrez, Source | orosomucoid 2 | 18111 | -0.159 | -0.0016 | No |
| 90 | IL17C | NA | IL17C Entrez, Source | interleukin 17C | 18295 | -0.172 | -0.0067 | No |
| 91 | PLA2G7 | NA | PLA2G7 Entrez, Source | phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) | 18654 | -0.197 | -0.0181 | No |
| 92 | PLA2G2D | NA | PLA2G2D Entrez, Source | phospholipase A2, group IID | 18866 | -0.215 | -0.0238 | No |
| 93 | CCL3 | NA | CCL3 Entrez, Source | chemokine (C-C motif) ligand 3 | 18961 | -0.223 | -0.0252 | No |
| 94 | ADORA2A | NA | ADORA2A Entrez, Source | adenosine A2a receptor | 18962 | -0.223 | -0.0232 | No |
| 95 | S100A9 | NA | S100A9 Entrez, Source | S100 calcium binding protein A9 | 19218 | -0.246 | -0.0303 | No |
| 96 | PLA2G2E | NA | PLA2G2E Entrez, Source | phospholipase A2, group IIE | 19411 | -0.261 | -0.0350 | No |
| 97 | CCL5 | NA | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 19429 | -0.263 | -0.0332 | No |
| 98 | HDAC7 | NA | 19453 | -0.265 | -0.0316 | No | ||
| 99 | GHSR | NA | GHSR Entrez, Source | growth hormone secretagogue receptor | 19478 | -0.268 | -0.0300 | No |
| 100 | NFRKB | NA | NFRKB Entrez, Source | nuclear factor related to kappaB binding protein | 19558 | -0.276 | -0.0303 | No |
| 101 | CYBB | NA | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 20252 | -0.349 | -0.0526 | No |
| 102 | IL18RAP | NA | IL18RAP Entrez, Source | interleukin 18 receptor accessory protein | 20475 | -0.377 | -0.0573 | No |
| 103 | GPR68 | NA | GPR68 Entrez, Source | G protein-coupled receptor 68 | 20646 | -0.398 | -0.0598 | No |
| 104 | CCL13 | NA | CCL13 Entrez, Source | chemokine (C-C motif) ligand 13 | 20652 | -0.399 | -0.0563 | No |
| 105 | ABCF1 | NA | ABCF1 Entrez, Source | ATP-binding cassette, sub-family F (GCN20), member 1 | 20706 | -0.407 | -0.0545 | No |
| 106 | NFKB1 | NA | NFKB1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (p105) | 21956 | -0.621 | -0.0947 | No |
| 107 | CCL11 | NA | CCL11 Entrez, Source | chemokine (C-C motif) ligand 11 | 21975 | -0.625 | -0.0896 | No |
| 108 | IL1A | NA | IL1A Entrez, Source | interleukin 1, alpha | 22157 | -0.658 | -0.0902 | No |
| 109 | AOC3 | NA | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 22222 | -0.669 | -0.0864 | No |
| 110 | IRAK2 | NA | IRAK2 Entrez, Source | interleukin-1 receptor-associated kinase 2 | 22634 | -0.761 | -0.0945 | No |
| 111 | PRDX5 | NA | PRDX5 Entrez, Source | peroxiredoxin 5 | 23262 | -0.894 | -0.1093 | No |
| 112 | LTB4R | NA | LTB4R Entrez, Source | leukotriene B4 receptor | 23488 | -0.948 | -0.1088 | No |
| 113 | RELA | NA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 23495 | -0.950 | -0.1003 | No |
| 114 | NLRP3 | NA | 24509 | -1.245 | -0.1260 | No | ||
| 115 | AIMP1 | NA | 24553 | -1.262 | -0.1160 | No | ||
| 116 | TGFB1 | NA | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 24608 | -1.278 | -0.1062 | No |
| 117 | RIPK2 | NA | RIPK2 Entrez, Source | receptor-interacting serine-threonine kinase 2 | 24745 | -1.325 | -0.0989 | No |
| 118 | CXCL2 | NA | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 24873 | -1.368 | -0.0910 | No |
| 119 | CEBPB | NA | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 25505 | -1.643 | -0.0990 | No |
| 120 | F11R | NA | F11R Entrez, Source | F11 receptor | 25611 | -1.693 | -0.0873 | No |
| 121 | NFX1 | NA | NFX1 Entrez, Source | nuclear transcription factor, X-box binding 1 | 25804 | -1.799 | -0.0777 | No |
| 122 | CXCL1 | NA | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 25823 | -1.806 | -0.0617 | No |
| 123 | AOX1 | NA | AOX1 Entrez, Source | aldehyde oxidase 1 | 25963 | -1.893 | -0.0494 | No |
| 124 | SIGIRR | NA | SIGIRR Entrez, Source | single immunoglobulin and toll-interleukin 1 receptor (TIR) domain | 26517 | -2.313 | -0.0484 | No |
| 125 | PARP4 | NA | PARP4 Entrez, Source | poly (ADP-ribose) polymerase family, member 4 | 26694 | -2.491 | -0.0319 | No |
| 126 | NMI | NA | NMI Entrez, Source | N-myc (and STAT) interactor | 26969 | -2.946 | -0.0148 | No |
| 127 | CD40 | NA | CD40 Entrez, Source | CD40 molecule, TNF receptor superfamily member 5 | 26999 | -3.035 | 0.0121 | No |